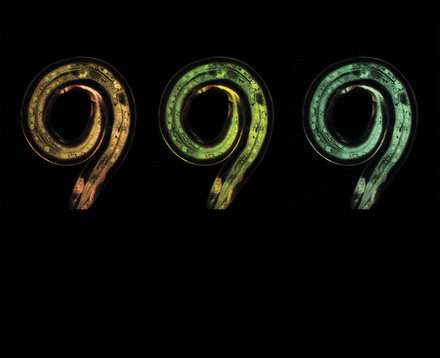Do you have results from a mutant screen to publish? G3’s Mutant Screen Reports allow you to publish succinct descriptions of useful genetic screens in a convenient format. The Reports fulfill one of G3’s goals: to make data from screens available to the community in a timely fashion.
If you gently touch the front half of a Caenorhabditis elegans nematode with an eyebrow hair glued to a toothpick, it will stop and reverse course. Eyelashes will do the trick too, apparently, though it hurts more for the experimenter/hair donor to sacrifice them. Over several decades, this simple and quantifiable behavioral response has proven a powerful model for understanding mechanosensation—a crucial component of hearing, touch, the sense of balance, and other important nervous system functions—as well as the development of specialized neuron subtypes.
The “gentle touch response,” as it is known, is separate from reactions to other touch stimuli—such as being jabbed by a wire or cosying up to a mate—and is mediated by six touch receptor neurons coupled to the animal’s cuticle (its “skin”). Genes needed for the development and function of these specialized neurons have been identified by several mutagenesis and gene expression studies.
However, one major type of gene has mostly escaped detection by mutagenesis screens: genes with such central functions to the animal that mutating or knocking them down causes paralysis, sterility, or death. Such pleiotropic genes are routinely missed by standard screens because there’s no way to assay neuronal phenotypes in a dead or paralyzed worm. So how can we identify pleiotropic genes needed for touch sensitivity?
Several years ago, Marty Chalfie’s lab developed a C. elegans strain that showed enhanced effects of RNA interference (RNAi) in its neurons, and a dampened RNAi response in other cell types. Using this strain in feeding RNAi screens allows experimenters to target genes with non-viable phenotypes and to observe their selective effects on the nervous system.
In the latest issue of G3, the lab turned to this neuronally enhanced RNAi approach to systematically identify pleiotropic genes affecting gentle touch sensitivity. Chen et al. found 61 genes that show reduced touch sensitivity when targeted by neuronally enhanced RNAi, out of 849 genes normally required for viability or movement in standard RNAi experiments. These included genes involved in cellular housekeeping functions, transcriptional regulation, protein degradation, cellular adhesion, cytoskeletal structure and function, endo/exocytosis, mitochondrial function, calcium signaling, and other signaling cascades.
The team tested 18 of the hits by examining the phenotypes of corresponding loss-of-function alleles. All 18 showed some degree of touch insensitivity, demonstrating the screen had a low false-positive rate.
One of the confirmed genes was homologous to the human gene TXNDC9, which is involved in microtubule growth and organization. Since touch receptor neurons require a unique type of microtubule filament for their function, the team followed up on this candidate gene (which they named txdc-9). The effects of txdc-9 on touch sensitivity were cell autonomous, and txdc-9 was needed for tubulin acetylation and microtubule function in the touch receptor neurons.
More work is needed to follow up on other hits from the screen, but the results show that neuronally enhanced RNAi screens can effectively complement standard mutagenesis for studies of neuronal phenotypes, opening up new and fruitful lines of inquiry even in well-studied systems. So C. elegans geneticists may need to keep their eyebrow tweezers handy for some time yet.
Don’t hide away useful mutant screen results in your lab books; submit a Mutant Screen Report today using our template, and publish your findings sooner!
CITATION
Chen X., M. D. Cuadros, & M. Chalfie (2015). Identification of Nonviable Genes Affecting Touch Sensitivity in Caenorhabditis elegans Using Neuronally Enhanced Feeding RNA Interference, G3, 5 (3) 467-475. DOI: http://dx.doi.org/10.1534/g3.114.015776
http://www.g3journal.org/content/5/3/467.full